Expandiendo las fronteras de la Ciencia
26 Mar 2024
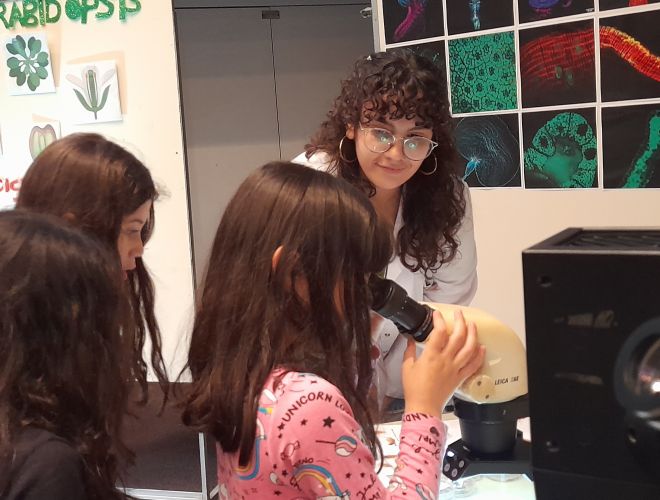
Sumate a la edición 2024 de “¿Qué Hacemos en el Instituto Leloir?”, nuestras jornadas destinadas a que conozcas qué y cómo investigamos
Si estás buscando un lugar para iniciar o continuar tu carrera científica, no te pierdas nuestro ya tradicional “QHL”, que durante dos días ofrece charlas (virtuales y/o presenciales) en las que nuestros investigadores te contarán los temas que estudiamos y qué oportunidades hay para que te unas a alguno de nuestros grupos. También tendrás la posibilidad de visitar el área de Microscopía.
Ver más
25 Mar 2024
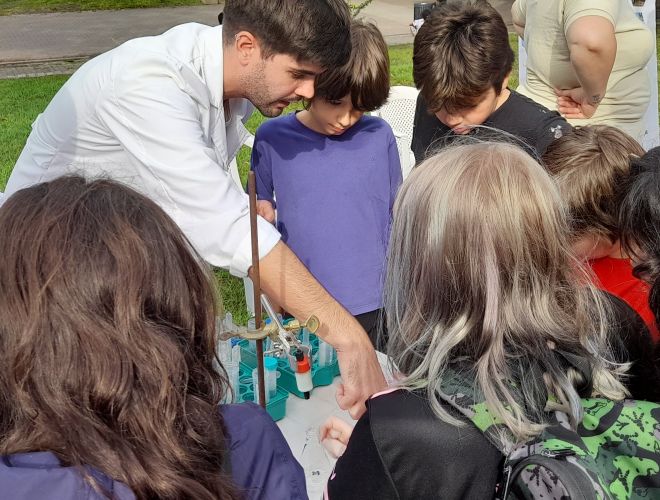
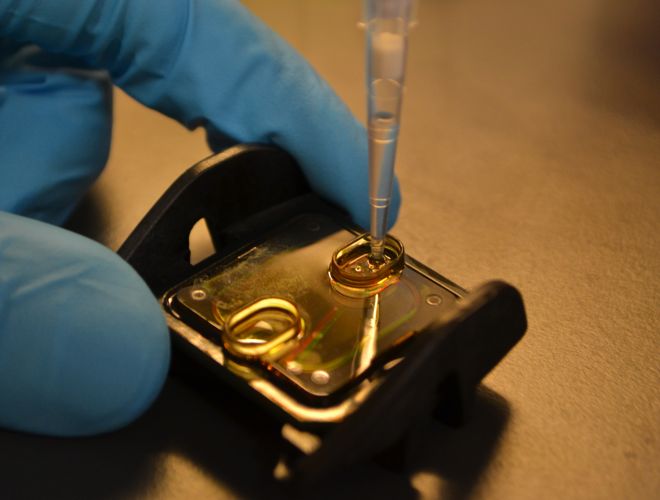
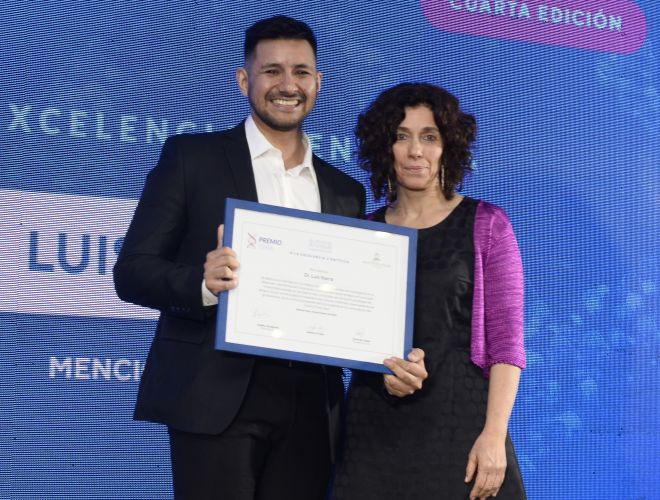
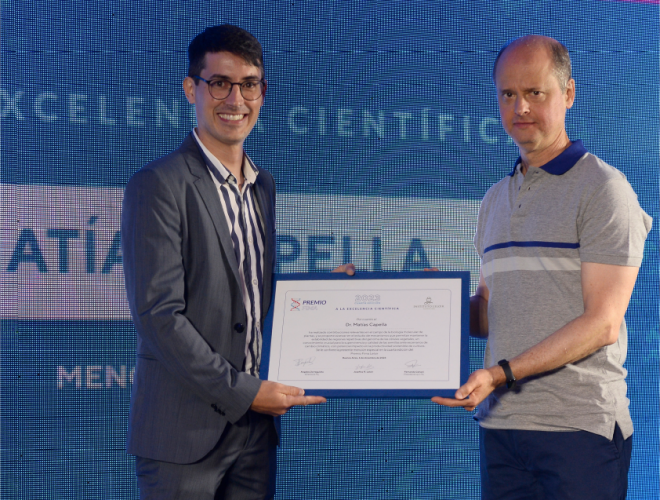
Un investigador de nuestro Instituto, detrás del éxito de las escuelas argentinas en el Concurso Internacional de Crecimiento de Cristales
Se trata del doctor en Química Sebastián Klinke, también presidente de la Asociación Argentina de Cristalografía, organización que desde 2014 impulsa cursos de capacitación sobre la disciplina a docentes de todo el país. En el último certamen mundial de Cristalografía, trabajos de dos escuelas patagónicas obtuvieron el primero y el tercer puesto.
Ver más
Áreas de investigación