Expandiendo las fronteras de la Ciencia
09 Abr 2024
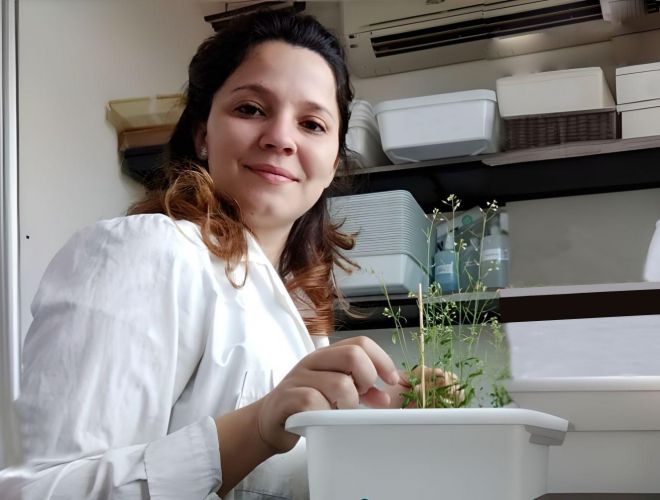
Una joven investigadora de nuestro Instituto ganó el premio a mejor póster en un congreso internacional
Se trata de la bióloga Caterina Sister, becaria de doctorado del CONICET en nuestro Laboratorio de Genética del Desarrollo Neural. El reconocimiento lo recibió durante el último encuentro de la Sociedad Latinoamericana de Biología del Desarrollo (LASDB).
Ver más
26 Mar 2024
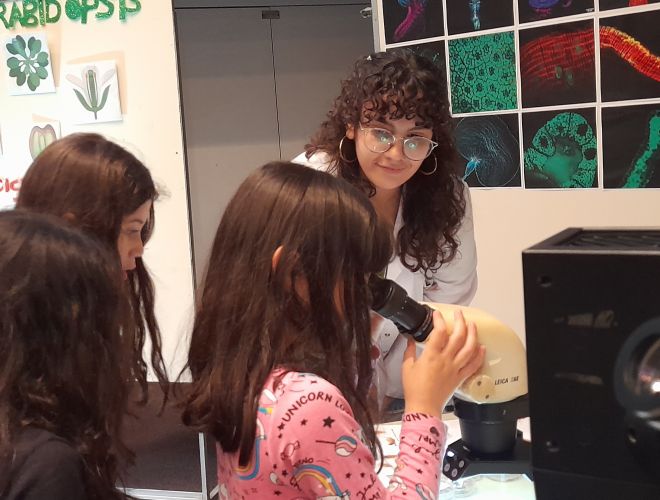
Sumate a la edición 2024 de “¿Qué Hacemos en el Instituto Leloir?”, nuestras jornadas destinadas a que conozcas qué y cómo investigamos
Si estás buscando un lugar para iniciar o continuar tu carrera científica, no te pierdas nuestro ya tradicional “QHL”, que durante dos días ofrece charlas (virtuales y/o presenciales) en las que nuestros investigadores te contarán los temas que estudiamos y qué oportunidades hay para que te unas a alguno de nuestros grupos. También tendrás la posibilidad de visitar el área de Microscopía.
Ver más
Áreas de investigación